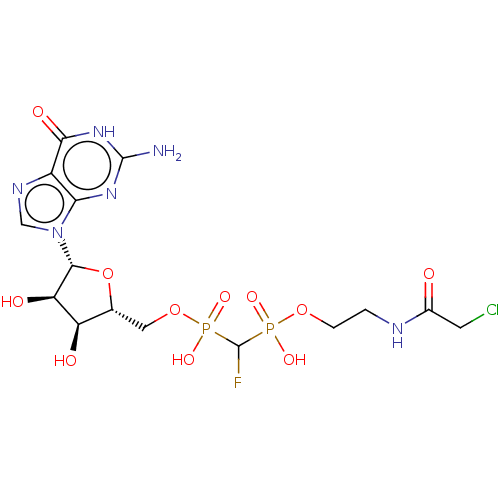
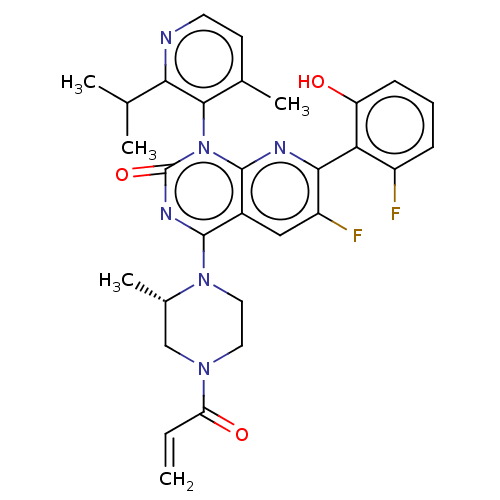
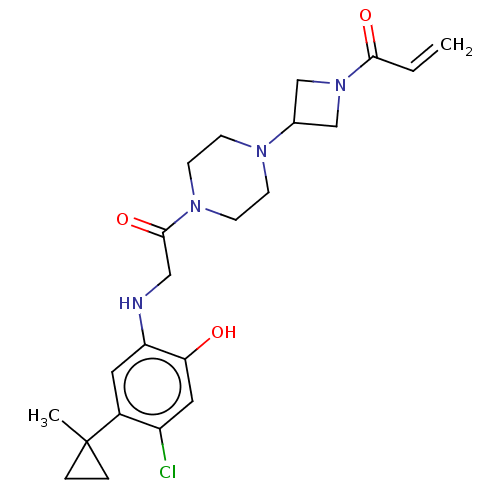
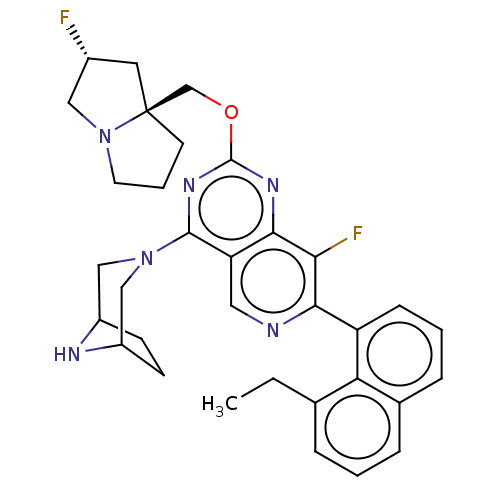
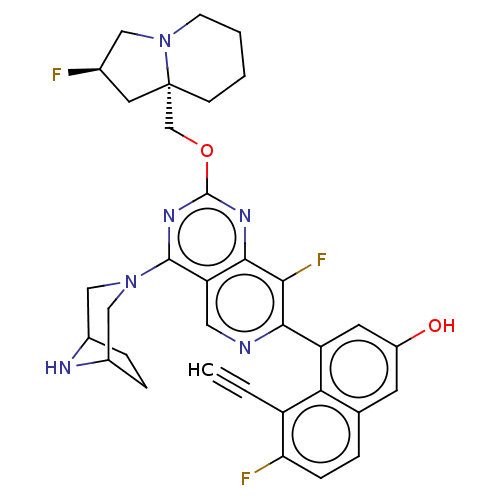
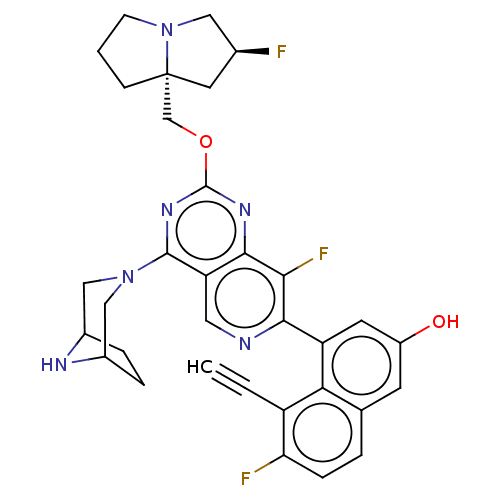
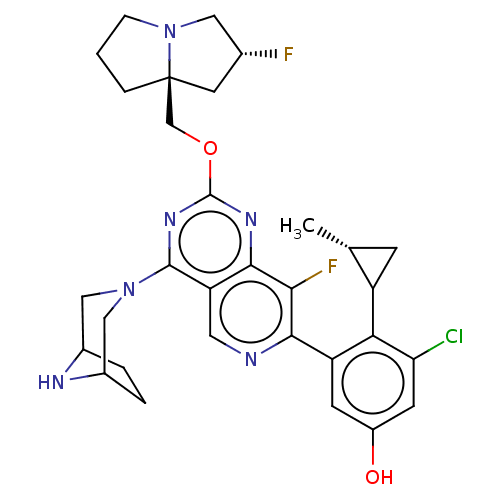
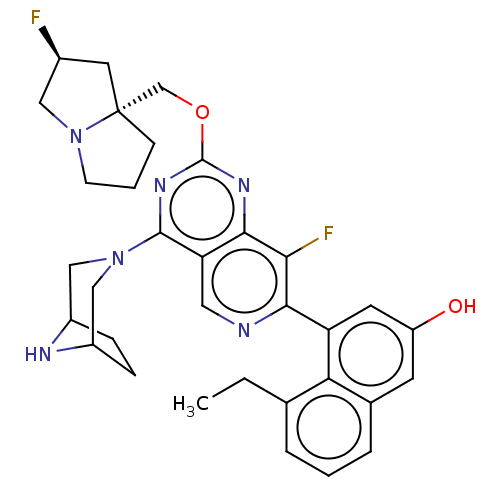
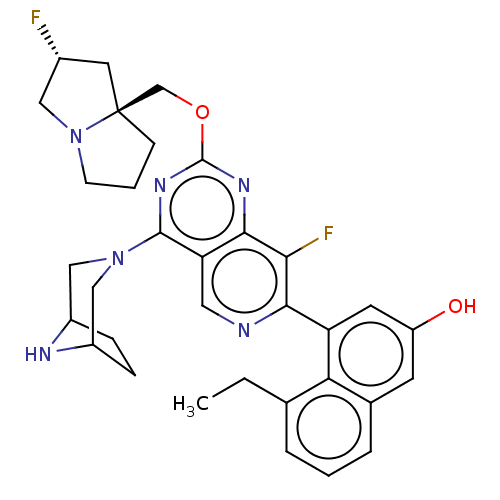
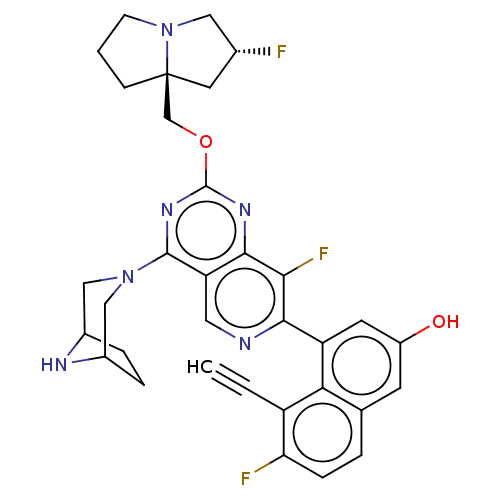
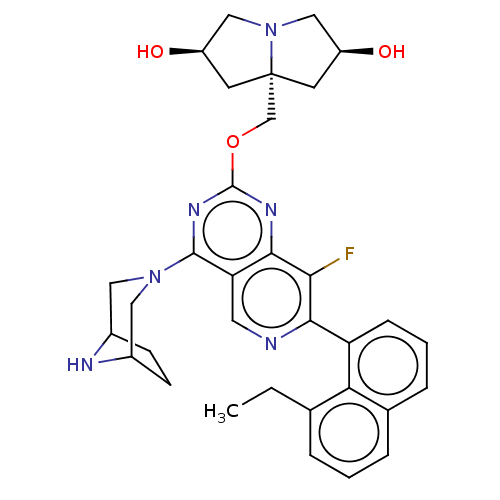
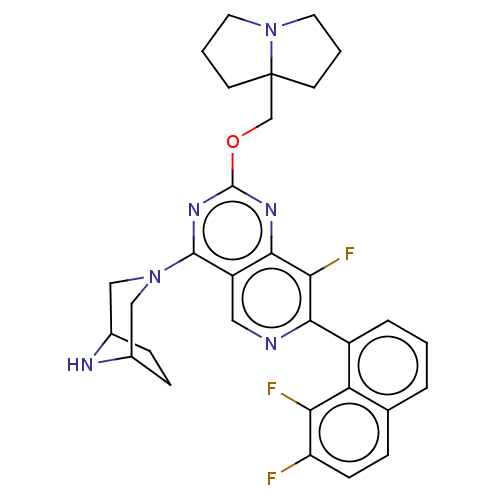
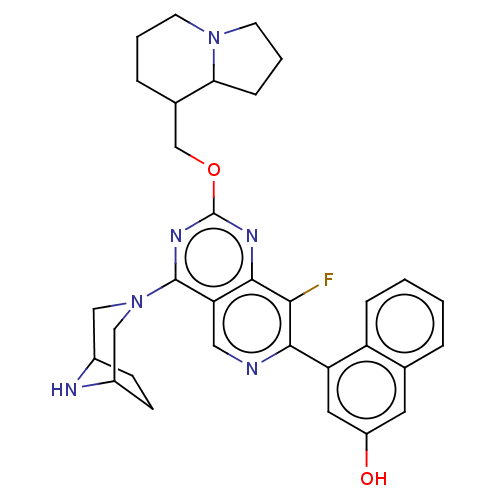
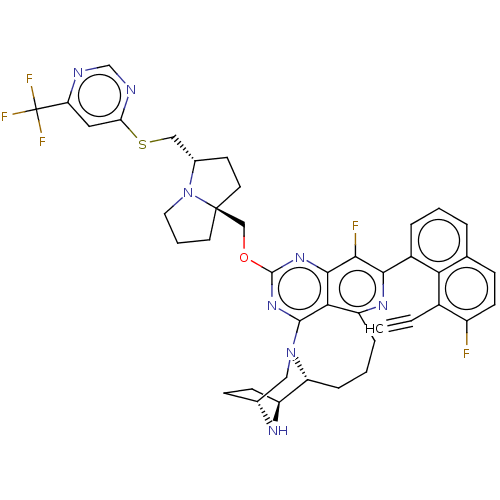
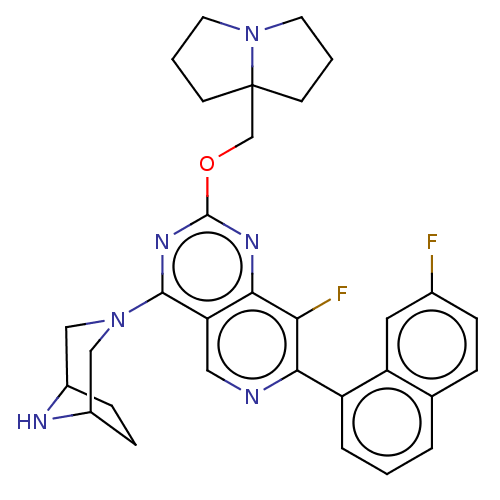
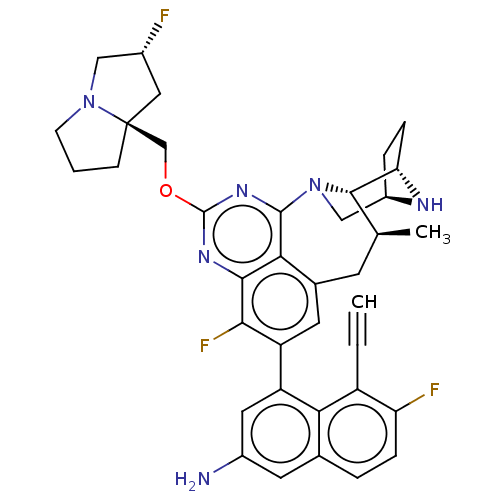
Affinity DataKi: 9nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 280nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 374nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 380nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 382nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 920nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.60E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.20E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.50E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 2.70E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.50E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.70E+3nMAssay Description:Inhibition of recombinant KRAS G12C mutant (unknown origin) assessed as rate of inactivation by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 3.70E+3nMAssay Description:Binding affinity to KRAS (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 7.50E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 8.00E+3nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+3nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: >1.00E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 1.22E+4nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: 1.52E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: >2.00E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 2.41E+4nMAssay Description:Binding affinity to KRAS G12C mutant (unknown origin) assessed as inhibition constant incubated for 24 hrs by LC-MS analysisMore data for this Ligand-Target Pair
Affinity DataKi: >2.50E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 3.10E+4nMAssay Description:Binding affinity to KRAS (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 3.60E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: 5.00E+4nMAssay Description:Binding affinity to GMP-stabilized FLAG-tagged KRAS G12C mutant (unknown origin) using desthiobiotin-GTP probe by alphascreen assayMore data for this Ligand-Target Pair
Affinity DataKi: >6.40E+4nMAssay Description:Binding affinity to hexahistidine-tagged recombinant human KRas G12C mutant expressed in Escherichia coli BL21 (DE3) in presence of GDPMore data for this Ligand-Target Pair
Affinity DataKi: 8.60E+4nMAssay Description:Binding affinity to KRAS (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataKi: 2.00E+5nMAssay Description:Inhibition of KRAS G12C mutant (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+5nMAssay Description:Inhibition of KRAS G12C mutant (unknown origin)More data for this Ligand-Target Pair
Affinity DataKi: 2.00E+5nMAssay Description:Binding affinity to KRAS (unknown origin) assessed as inhibition constantMore data for this Ligand-Target Pair
Affinity DataIC50: 0.00100nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: <0.00200nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00600nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: 0.00700nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0120nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0200nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0270nMAssay Description:This Example illustrates that exemplary compounds of the present invention inhibit the phosphorylation of ERK downstream of KRAS G12D. AGS cells (ATC...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0400nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Compounds were tested for binding to GDP-loaded KRAS G12D in a 384-well assay format using a TR-FRET probe displacement assay in buffer consisting of...More data for this Ligand-Target Pair
Affinity DataIC50: 0.0500nMAssay Description:Biochemical compound potencies are assessed by evaluating inhibition of SOS1I-mediated nucleotide exchange in KRAS G121D. In this assay, the SOS1-pro...More data for this Ligand-Target Pair



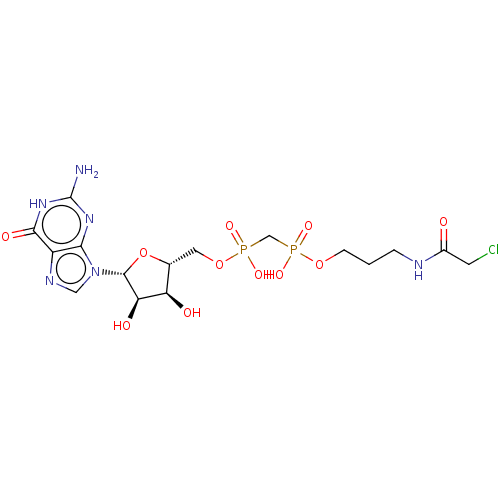


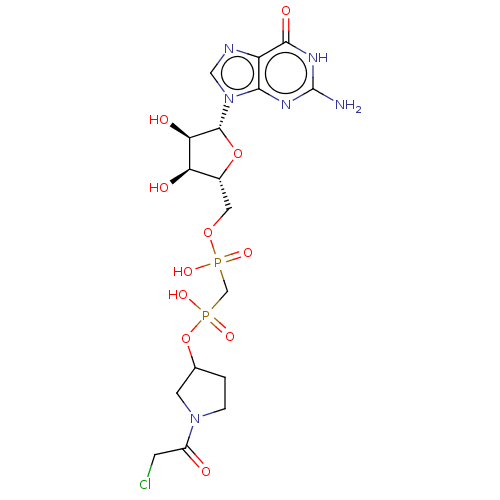
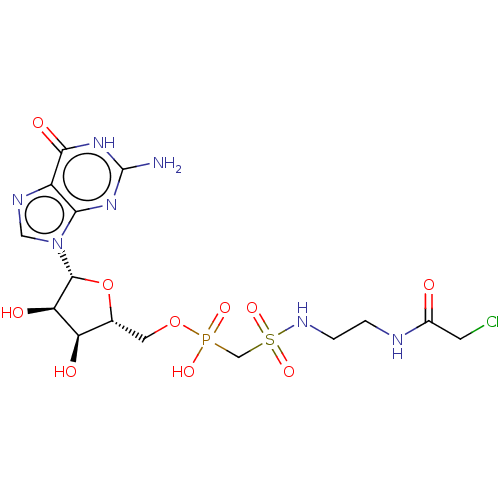


 3D Structure (crystal)
3D Structure (crystal)